
- On 11 Apr 2023
- In Computer Science
- Tags: Tips and Tricks, Python, EDA
Reduce Size Constraints by Addressing Data Types
Every dataset we use in the world has some sort of size constraint that comes with it. How do we take the data we use in one project or system and make it more useable for downstream analysis in order to support data pipelines and machine learning operations? This work can start in many places, but it is essential to include the reclassification of columns related to variable types as early in the process as possible. If you are familiar with pandas, you probably think of this as setting the Dtypes.
After creating a pandas DataFrame, the next step should always be to look at the Dtype of each column. If there is a more appropriate Dtype for any of the columns, they should all be converted before any subsequent coding or data analysis is done. The most important reason for doing this at the very beginning is that the potential for saving memory is huge.
Why do we care about saving a huge amount of data?
It all comes down to the speed and efficiency of which your data is loaded, processed, transformed, analyzed, and visualized. Our work at Foretify involves many types of datasets, including, geospatial data and machine learning models. Like we mentioned earlier, any meaningful dataset is going to be large. When you combine multiple large datasets, the results are a gargantuan dataset which will inevitably increase the amount of time it takes to process. By setting up the data to be as small as possible from the beginning, it makes coding, analyzing, and processing much easier and more efficient down the code road.
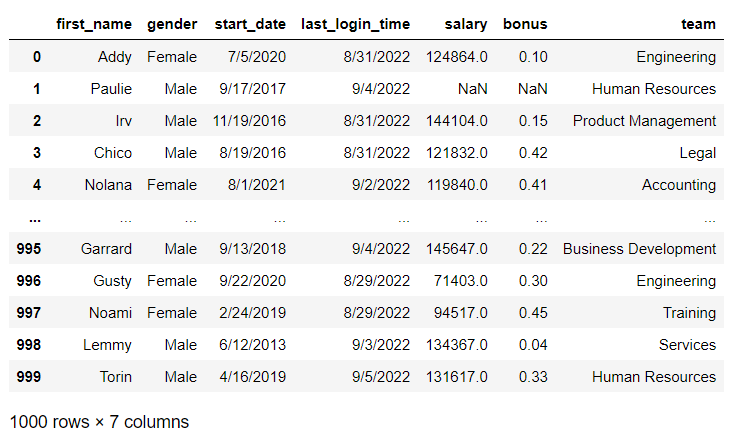
To illustrate how to convert Dtypes in a DataFrame, I created a mock dataset using a site called Mockaroo. The dataset contains fictional employee records including name, gender, start date, last login time, salary, bonus, and team assignment. As shown below, once the csv has been read into a DataFrame, the .info() method can be used to see a quick summary of the data showing columns, Dtypes, number of non-null values, and the rangeindex provided for the index axis:
import pandas as pd
df = pd.read_csv('mock_employee_data.csv')
df
df.info()
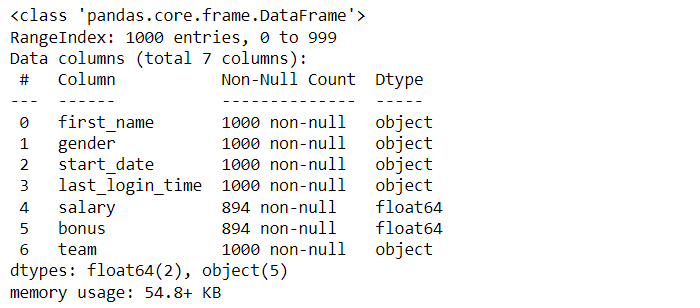
When data is first read into a pandas DataFrame, the textual data is automatically created as an object (or string) Dtype and the numerical data is created as a float, even the data only includes whole numbers.
Each of the columns is automatically created as an object or a float, and the memory is 54.8 KB. However, many of the columns can be converted to a more appropriate Dtype:
- Gender and team can both be a category Dtype
- Start date and Last Login Time can both be converted to the datetime Dtype
- Salary can be converted to the integer Dtype
Check for Null Values
The .astype() and and pd.to_datetime() methods can be used to convert them all to their new Dtypes, but this requires that no null values are present in any of the numerical columns being converted. If there are any missing or NaN values, the .astype() method will result in an error because pandas cannot convert a missing value to an integer or a float.
As mentioned previously, the .info() output will display the number of non null values in a column. If the number of non null values is less than the length of your DataFrame, there are non null values in your dataset. Another way to check for null values is using the .hasnans method:
df['salary'].hasnans df['bonus'].hasnans
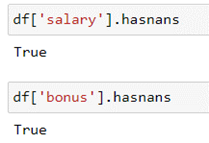
Fill Null Values
Since there are rows that contain null values, they will need to be resolved before we can proceed. There are several ways to address null values in a DataFrame, but for this article I’ve chosen to use the .fillna() method to demonstrate how changing the Dtype alone can decrease the memory usage of your dataset.
The column will be overwritten to replace the NaN values with 00 in order to retain as much of the data as possible.
df['salary'] = df['salary'].fillna(0) df['bonus'] = df['bonus'].fillna(0)
Double check the df to make sure there are no more NaNs:
df['salary'].hasnans df['bonus'].hasnans

Convert the Dtypes & See the Difference!
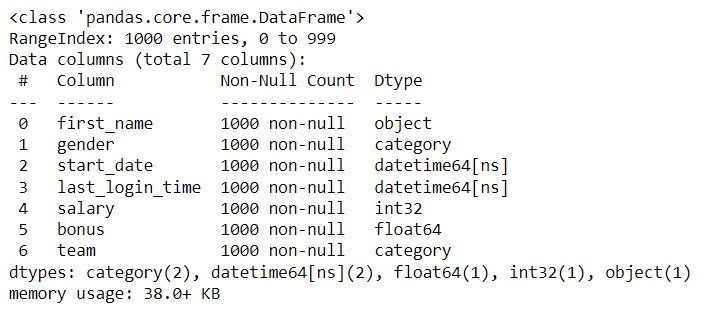
Now the columns are ready to be converted by overwriting each column using the .astype() and pd.to_datetime() methods. Without removing a single row from the dataset, the conversion resulted in the memory usage decreasing from 54.8 KB to 38.0 KB—that’s a memory savings of 30%! In a dataset as small as my sample (only one thousand rows), this doesn't make too much of an impact. But just imagine what a huge impact this could make on a data set with one million or more rows! This savings in memory will result in also saving precious seconds or even minutes of processing time.
df['gender'] = df['gender'].astype('category')
df['start_date'] = pd.to_datetime(df['start_date'])
df['last_login_time'] = pd.to_datetime(df['last_login_time'])
df['salary'] = df['salary'].astype('int')
df['team'] = df['team'].astype('category')
Join our mailing list!
Did you find what you need?
For more information or a product demonstration, please feel
free to reach out.
Contact us
Posted by Rebecca Beitel| 11 Apr 2023 |About Us
Rebecca Beitel is a GIS Systems Engineer with an MBA in Location Analytics. Her disciplined and quality-focused approach has helped her excel in building end-to-end solutions. She thrives on learning new concepts and technologies. When she's not exploring the latest advancements in the field, she can be found hiking, reading, or trying out new recipes in the kitchen.